-Search query
-Search result
Showing all 44 items for (author: prazak & v)
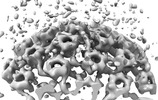
EMDB-18482: 
Herpes simplex virus 1 capsid (WT) vertices in perinuclear NEC-coated vesicles determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D

EMDB-18484: 
Herpes simplex virus 1 nuclear egress complex (WT) determined in situ from perinuclear vesicles
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D

EMDB-17974: 
Pseudorabies virus cytosolic C-capsid (US3 KO) vertices determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D

EMDB-17975: 
Pseudorabies virus primary enveloped (perinuclear) C-capsid (US3 KO) vertices determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D

EMDB-17976: 
Pseudorabies nuclear C-capsids (US3 KO) vertices determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D

EMDB-18473: 
Subtomogram average of pseudorabies virus nuclear egress complex helical form (UL31/34) determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D

EMDB-18474: 
Subtomogram average of pseudorabies virus nuclear egress complex (UL31/34) determined in situ
Method: subtomogram averaging / : Prazak V, Grange M, Vasishtan D
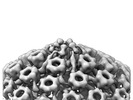

EMDB-18479: 
Pseudorabies virus cytosolic C-capsid (WT) vertices determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D

EMDB-18480: 
Pseudorabies virus nuclear C-capsid (WT) vertices determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D
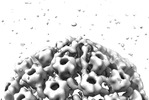

EMDB-18481: 
Herpes simplex virus 1 cytosolic C-capsid (WT) vertices determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D
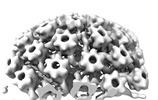
EMDB-18483: 
Herpes simplex virus 1 nuclear C-capsid (WT) vertices determined in situ
Method: subtomogram averaging / : Mironova Y, Prazak V, Vasishtan D
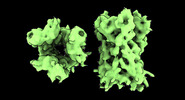
EMDB-17704: 
Subtomogram average of Vaccinia A10 trimer with open center from in vitro cores
Method: subtomogram averaging / : Turonova B, Liu J
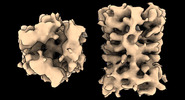
EMDB-17708: 
Subtomogram average of Vaccinia A10 trimer with tight center from in vitro cores
Method: subtomogram averaging / : Turonova B, Liu J

EMDB-17753: 
Subtomogram average of Vaccinia A10 trimer from in situ cores
Method: subtomogram averaging / : Turonova B, Liu J
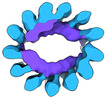

EMDB-15532: 
Plasmodium falciparum sporozoite subpellicular microtubule with interrupted luminal helix determined in situ
Method: subtomogram averaging / : Prazak V, Ferreira JL, Vasishtan D, Grunewald K, Siggel M, Hentzschel F, Pietsch E, Kosinski J, Frischknecht F, Gilberger TW

EMDB-15534: 
Plasmodium falciparum gametocyte subpellicular microtubule with 13 protofilaments determined in situ
Method: subtomogram averaging / : Ferreira JL, Prazak V, Vasishtan D, Grunewald K, Siggel M, Hentzschel F, Pietsch E, Kosinski J, Frischknecht F, Gilberger TW

EMDB-15535: 
Plasmodium falciparum gametocyte subpellicular microtubule with 14 protofilaments determined in situ
Method: subtomogram averaging / : Ferreira JL, Prazak V, Vasishtan D, Grunewald K, Siggel M, Hentzschel F, Pietsch E, Kosinski J, Frischknecht F, Gilberger TW

EMDB-15536: 
Plasmodium falciparum gametocyte subpellicular microtubule with 15 protofilaments determined in situ
Method: subtomogram averaging / : Ferreira JL, Prazak V, Vasishtan D, Grunewald K, Siggel M, Hentzschel F, Pietsch E, Kosinski J, Frischknecht F, Gilberger TW

EMDB-15537: 
Plasmodium falciparum gametocyte subpellicular microtubule with 16 protofilaments determined in situ
Method: subtomogram averaging / : Ferreira JL, Prazak V, Vasishtan D, Grunewald K, Siggel M, Hentzschel F, Pietsch E, Kosinski J, Frischknecht F, Gilberger TW

EMDB-15538: 
Plasmodium falciparum gametocyte subpellicular microtubule with 17 protofilaments determined in situ
Method: subtomogram averaging / : Ferreira JL, Prazak V, Vasishtan D, Grunewald K, Siggel M, Hentzschel F, Pietsch E, Kosinski J, Frischknecht F, Gilberger TW

EMDB-15539: 
Plasmodium falciparum gametocyte subpellicular microtubule with 18 protofilaments determined in situ
Method: subtomogram averaging / : Ferreira JL, Prazak V, Vasishtan D, Grunewald K, Siggel M, Hentzschel F, Pietsch E, Kosinski J, Frischknecht F, Gilberger TW

EMDB-15627: 
Tomogram of the Mimivirus genomic fiber
Method: electron tomography / : Villalta A, Schmitt A, Estrozi L, Quemin ERK, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Gruenewald K, Abergel C

EMDB-15628: 
Tomogram of the Mimivirus genomic fiber
Method: electron tomography / : Villalta A, Schmitt A, Estrozi L, Quemin ERK, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Gruenewald K, Abergel C

EMDB-15629: 
Tomogram of the Mimivirus genomic fiber
Method: electron tomography / : Villalta A, Schmitt A, Estrozi L, Quemin ERK, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Gruenewald K, Abergel C

EMDB-15630: 
Tomogram of the Mimivirus genomic fiber
Method: electron tomography / : Villalta A, Schmitt A, Estrozi L, Quemin ERK, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Gruenewald K, Abergel C

EMDB-13641: 
Structure of the Mimivirus genomic fibre asymmetric unit
Method: single particle / : Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
EMDB-14353: 
Structure of the Mimivirus genomic fibre in its compact 6-start helix form
Method: helical / : Villalta A, Schmitt A

EMDB-14354: 
Structure of the Mimivirus genomic fibre in its compact 5-start helix form
Method: helical / : Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C

EMDB-14355: 
Structure of the Mimivirus genomic fibre in its relaxed 5-start helix form
Method: helical / : Villalta A, Schmitt A

PDB-7ptv: 
Structure of the Mimivirus genomic fibre asymmetric unit
Method: single particle / : Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C

PDB-7yx3: 
Structure of the Mimivirus genomic fibre in its compact 6-start helix form
Method: helical / : Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C

PDB-7yx4: 
Structure of the Mimivirus genomic fibre in its compact 5-start helix form
Method: helical / : Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C

PDB-7yx5: 
Structure of the Mimivirus genomic fibre in its relaxed 5-start helix form
Method: helical / : Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C

EMDB-12188: 
Structure of DNA origami signpost from cryo-ET and subvolume averaging
Method: subtomogram averaging / : Silvester E, Vollmer B

PDB-7bho: 
DNA origami signpost designed model
Method: subtomogram averaging / : Silvester E, Vollmer B, Prazak V, Vasishtan D, Machala EA, Whittle C, Black S, Bath J, Turberfield AJ, Gruenewald K, Baker LA

EMDB-11123: 
Pre-fusion conformation of glycoprotein B of Herpes simplex virus 1
Method: subtomogram averaging / : Vollmer B, Prazak V, Vasishtan D, Jefferys EE, Hernandez-Duran A, Vallbracht M, Klupp B, Mettenleiter TC, Backovic M, Rey FA, Topf M, Gruenewald K

PDB-6z9m: 
Pseudoatomic model of the pre-fusion conformation of glycoprotein B of Herpes simplex virus 1
Method: subtomogram averaging / : Vollmer B, Prazak V, Vasishtan D, Jefferys EE, Hernandez-Duran A, Vallbracht M, Klupp B, Mettenleiter TC, Backovic M, Rey FA, Topf M, Gruenewald K

EMDB-10134: 
CryoEM structure of murine perforin-2 ectodomain in a pre-pore form
Method: single particle / : Ni T, Yu X, Gilbert RJC

EMDB-10135: 
CryoEM structure of murine perforin-2 ectodomain in a pore form
Method: single particle / : Ni T, Yu X, Gilbert RJC

EMDB-4995: 
Subtomogram average of a part of the rabies lyssavirus ribonucleoprotein
Method: subtomogram averaging / : Riedel C, Vasishtan D, Prazak V, Ghanem A, Conzelmann KK, Ruemenapf T

EMDB-4471: 
Cryo-SOFI enabling low-dose super-resolution correlative light and electron cryo-microscopy
Method: electron tomography / : Moser F, Prazak V, Mordhorst V, Andrade AM, Baker LA, Hagen C, Grunewald K, Kaufmann R

EMDB-4472: 
Cryo-SOFI enabling low-dose super-resolution correlative light and electron cryo-microscopy
Method: electron tomography / : Moser F, Prazak V, Mordhorst V, Andrade AM, Baker LA, Hagen C, Grunewald K, Kaufmann R

EMDB-4473: 
Cryo-SOFI enabling low-dose super-resolution correlative light and electron cryo-microscopy
Method: electron tomography / : Moser F, Prazak V, Mordhorst V, Andrade AM, Baker LA, Hagen C, Grunewald K, Kaufmann R

EMDB-3035: 
The structure of YnaI implies structural and mechanistic conservation in the MscS-family of mechanosensitive channels
Method: single particle / : Bottcher B, Prazak V, Rasmussen A, Black SS, Rasmussen T
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
